How to use the Literature-based manually-curated primer pairs databases?
To choose the database of interest please select one from a upper panel on the home page. Currently only database with primer pairs for antibiotic resistance genes amplification is available (LCPDb-ARG).
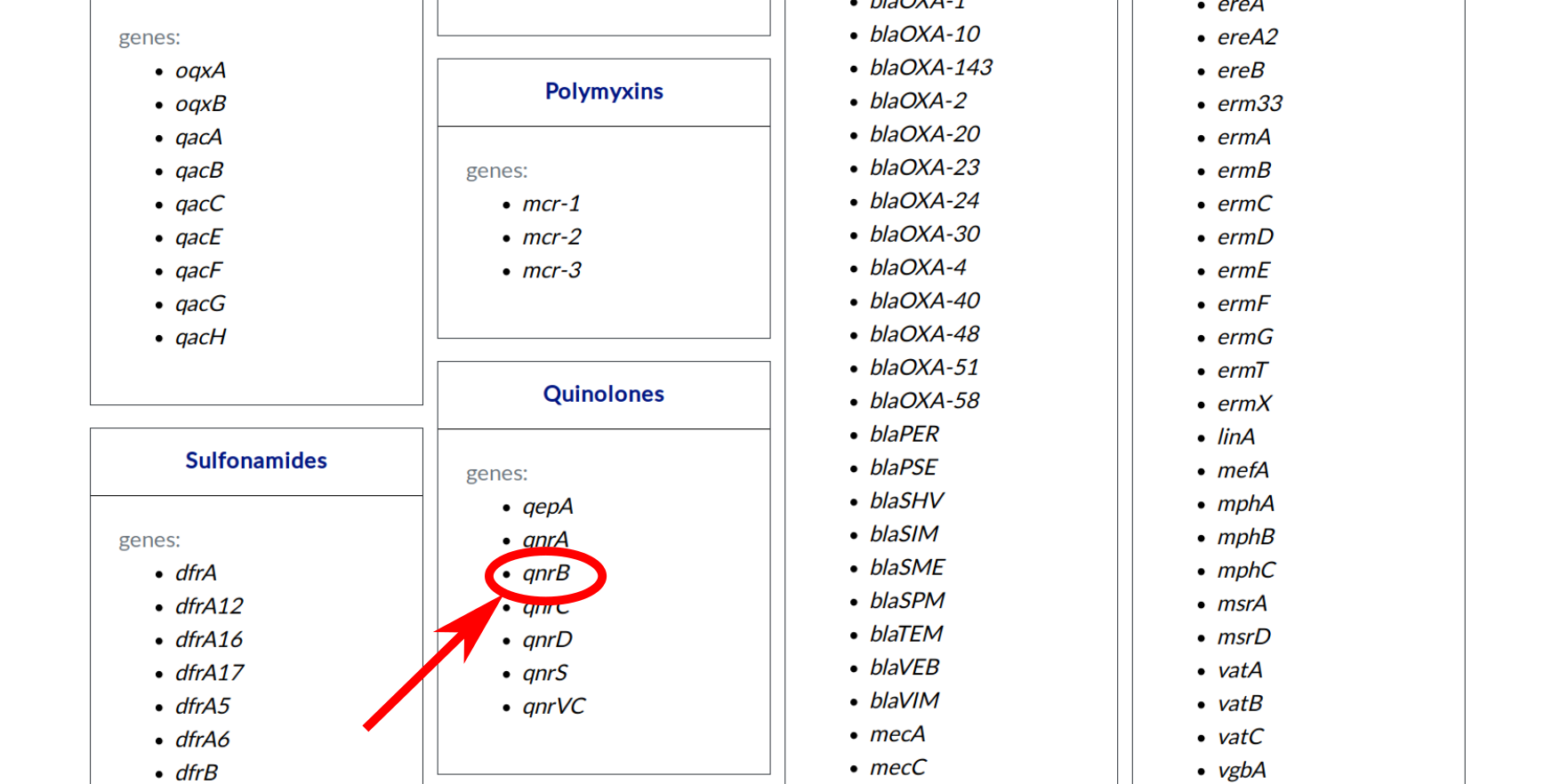
You are now on the page with all genes grouped by class of antibiotics for which primer pairs are available in LCPDb-ARG. To choose a gene please click one that you are interested in e.g. qnrB from quinolones family.
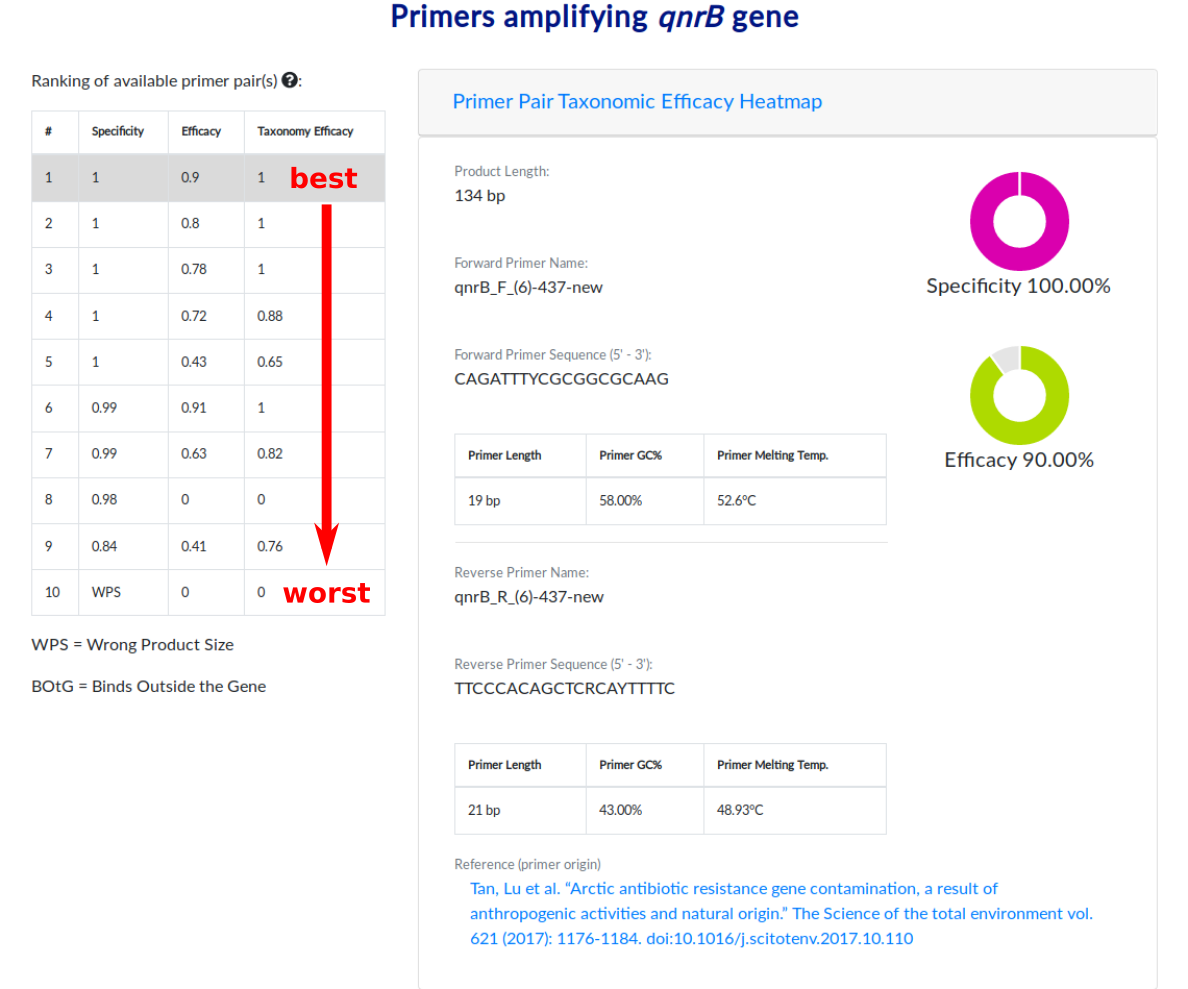
You are now on the page with the primer pairs ranking and information. On the left side you can see the ranking of PCR primer pairs dedicated for amplification of qnrB gene. Primer pairs specific for each antibiotic resistance gene were ranked based on their model success metric value (MSM). The value of MSM is the arithmetic average of parameters Specificity (S), Efficacy (E) and Taxonomic Efficacy (TE) normalized by the modified – in reference to the confusion matrix – accuracy parameter. The meanings of above-mentioned parameters are describe below:
- Specificity - If primer pair is specific to particular antibiotic resistant gene (1: very specific, 0: unspecific),
- Efficacy - How many variants of specific antibiotic resistance gene could be amplified (1: all variants, 0: none of variant),
- Taxonomic Efficacy - Within how many taxa specific ARG’s could be amplified with particular primer pair (1: all possible taxa, 0: none of taxon).
For general use we recommend to use primer pair from the first place in the ranking as that one is the best based on the model success metric value.
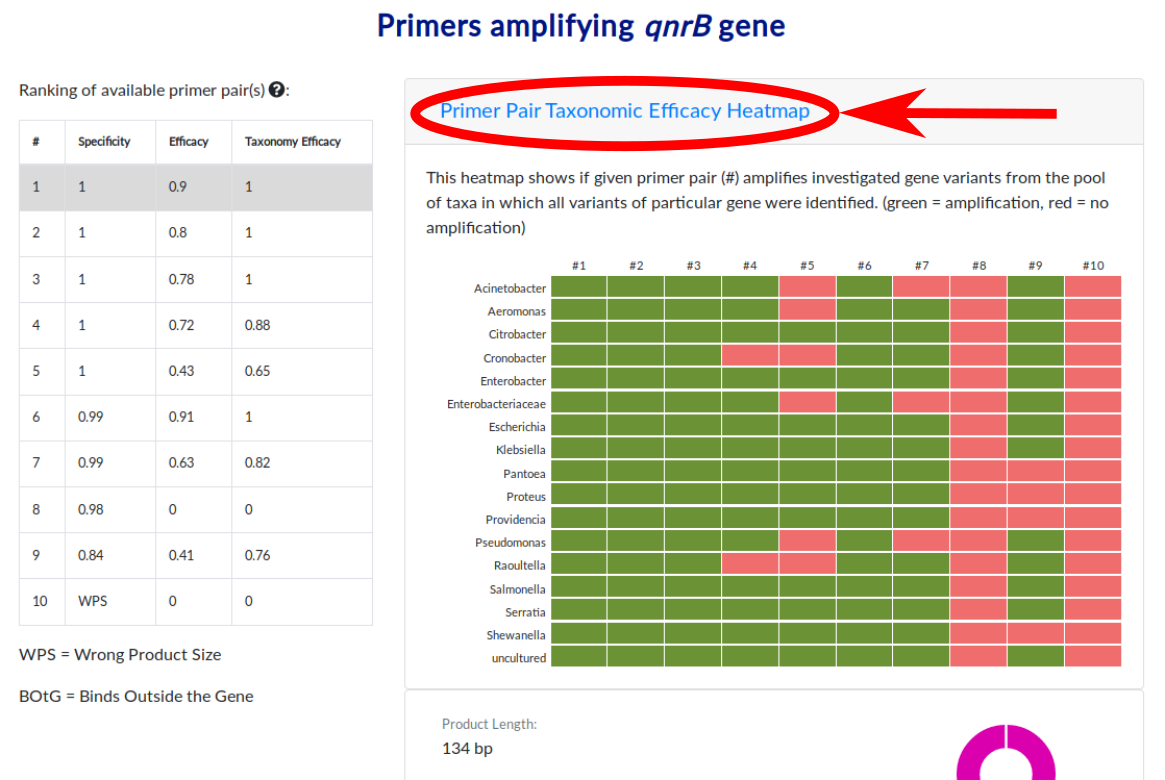
Additional feature of the database is the possibility to view into the heatmap showing taxon coverage of all primer pairs for particular ARG from the ranking (on the left side). To view the heatmap please click button "Primer Pair Taxonomic Efficacy Heatmap".